News
April, 2026
New preprint: measuring avidity with DNA origami
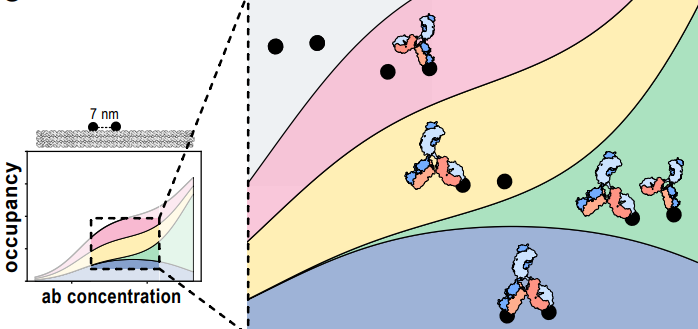
A new molecular programming preprint led by collaborators Iris Rocamonde Lago (currently postdoc in Erik Benson's group) and Björn Högberg. The work introduces a new method using DNA origami nanostructures and statistical physics / equilibrium thermodynamics to measure avidity — an elusive concept describing the binding strength of molecules connected by multiple points of contact. In this study, spatially separated antigens and their interaction with antibodies were used as a model system.
biorxiv.org preprint
April, 2026
VOLUMINEX annual meeting in Paris

The VOLUMINEX consortium met in Paris on 9–10 April 2026, bringing together all five partners at Sorbonne University for its annual meeting. The program combined technical updates across work packages with focused discussions on the emerging end-to-end pipeline and preparation for the upcoming project review. Alongside the science, the meeting provided time to align as a team and set direction for the next phase of the project. Photo shows us enjoying some French cuisine in central Paris.
VOLUMINEX in Paris for their 2026 annual meeting
April, 2026
Ian appointed to DDLS expert group for Cell and Molecular Biology

Ian Hoffecker has been appointed to the Data Driven Life Sciences (DDLS) expert group for Cell and Molecular Biology. Ian recently participated in the DDLS symposium on Data-Driven Systems Biology, held at Life City in Stockholm, bringing together researchers harnessing big data and systems approaches to decode complex biology.
DDLS Cell and Molecular Biology Research Area
DDLS Symposium: Data-Driven Systems Biology
March, 2026
Joel presents at FNANO in Munich
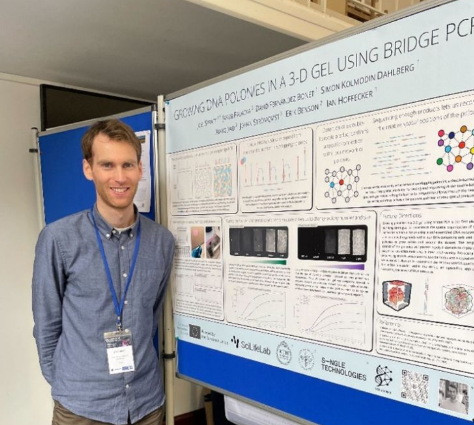

Last week, Molecular Programming group member and senior researcher Dr. Joel Spratt attended the Foundations of Nanoscience annual conference in Munich, Germany. On the 18th March, Joel presented his poster titled "Growing DNA Polonies in 3-D Gels Using Bridge PCR", which described the protocols developed in the VOLUMINEX project for bridge PCR inside 3-D gels.
The Foundations of Nanoscience conference is the top international conference for all things related to DNA nanotechnology, featuring the latest in nanostructure design and development from the field's biggest names. "It was great to bounce ideas around and see how our project fits into the bigger nanotech ecosystem" reported Joel. "And it was great to visit Munich as well!"
FNANO Conference Program
March, 2026
Ian appointed Digital Futures faculty member and participates in panel discussion

Ian has been appointed a Digital Futures faculty member, and recently participated in a panel discussion at the Digital Futures Young Scientist Conference (March 12th) together with Frank Jiang (KTH Innovation and FleetMQ) and Johanna Hård (Amun AI), and host Urban Forssell (KTH & Digital Futures). This lively discussion was about the challenges and opportunities of translating academic research into real-world applications, entrepreneurship, and interdisciplinary collaboration.
Digital Futures Young Scientist Conference recap
January, 2026
Happy new year, new paper, new people

Great start of 2026 - our new paper on measuring the spatial coherence of barcode networks is out. In this work, we present a theoretical framework for understanding how spatial information is encoded in molecular networks, and introduce a new metric, "spatial coherence", for quantifying the degree to which a network preserves spatial relationships. This metric can be used to evaluate and optimize sequencing-based microscopy techniques, or spatial networks more broadly. Congrats to lead author and PhD student David F. Bonet, together with Johanna, Shuai, Simon, and Ian!
And a warm welcome as well to KTH Masters students Johan Holmqvist (Physics) and Eskil Billsten (Applied mathematics) who are joining us for their thesis projects this sping.
Here's to a great year ahead!
cell.com/patterns/fulltext/S2666-3899(25)00276-4
November, 2025
Discovery of a hidden network
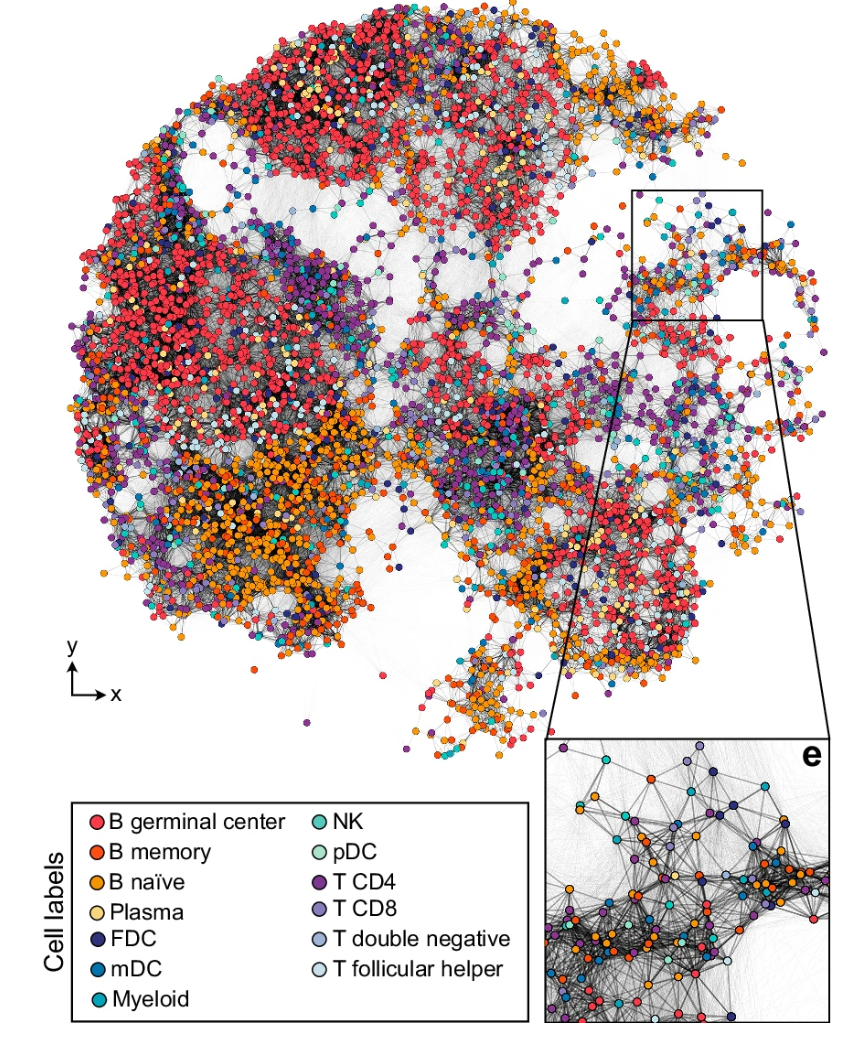
A new paper from our group is out! During a group journal club session, we made a serendipetous discovery of a hidden spatial network in data from the Slide-tags method developed by Fei Chen and colleagues at the Broad Institute. This network is formed as a consequence of the promiscuous diffusion of DNA barcodes from surface arrays to cell nuclei, and allowed us to perform surprisingly good spatial reconstructions using our algorithm "STRND" without the need for prior spatial reference mapping. Great effort by lead PhD student Simon K Dahlberg together with David Fernandez Bonet, and collaborators Lovisa Franzén and Patrik Ståhl. Well done!
nature.com/articles/s41467-025-65295-w
October, 2025
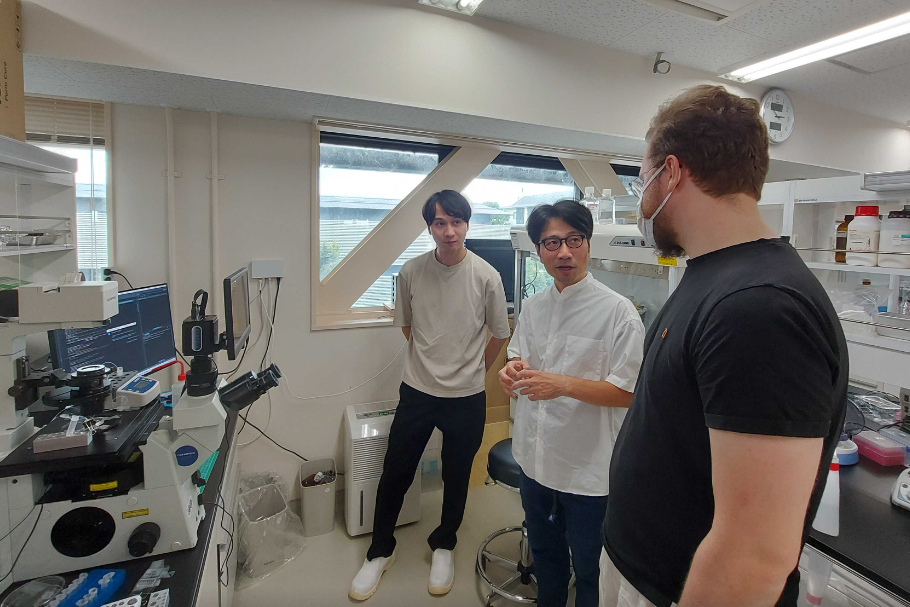
Autumn update
PhD student Simon K. Dahlberg begins a 3 month research exchange in Japan to help develop a super cool new sequencing-based spatial capture device in the high-tech lab of our collaborators Hirofumi Shintaku and Taikopaul Kaneko at the LiMe Institute, Kyoto University.
PhD student David and postdoc Anastassia attended FDN 2025 in Rome this year, and David's work on the promiscuity of spatial networks was selected for a talk. Well done! Also, we welcome spring Master's thesis student Sara Assariha who will be conducting an advanced-PCR DNA computing project together with mentor and senior researcher Joel Spratt. Welcome! Link to Shintaku lab

Aug 17, 2025
End of summer update
Back for more action after an eventful summer! Ian attended the Statphys Satellite meeting in Lviv, Ukraine (Statphys Lviv article) - building important connections and learning about the latest ideas in statistical physics, and Ian and Joel debuted our work on DNA-based data storage at the DBDS meeting in Prague this year.
Simon's paper on hidden networks was accepted to Nature Communications - congrats! And Voluminex collaborators Martin Weigt and Federico Scarpati visited Stockholm for a graph theory hackathon together with Ian and David.
Link to DBDS webpage

March 18, 2025
Voluminex project kickoff
Collaborators of the Voluminex project, coordinated by the Hoffecker group and KTH, arrive in Stockholm for the official project kickoff. In this 5 year international project, we will develop a network based imaging technology for 3D mapping of molecules in organoids.
Link to project webpage


Exciting announcement - our group at KTH/SciLifeLab will be coordinating a big cross-Europe project over the next 5 years to develop a 3D spatial molecular imaging technology from networks.
The project will be conducted together with partners Erik Benson at Karolinska/SciLifeLab Stockholm, Stockholm-based sequencing technology company Single Technologies AB led by CEO Johan Strömqvist, biologists Benedetta Artegiani and Delilah Hendriks at Princess Maxima Center for Pediatric Oncology in Utrecht, Netherlands, and physicist Martin Weigt at Sorbonne University in Paris.
We're funded by the EU through the EIC Pathfinder program. Read more about it here: News article link

November, 2024
Autumn update - talks, travel, and new members
This autumn, Ian, David, and Simon attended an exciting DNA30 conference in Baltimore where we learned about the latest breakthroughs in the molecular programming field. David and Simon presented the results of their recent research on finding hidden spatial information in DNA barcode networks.
We were also honored to host a talk here at SciLifeLab by Joshua Weinstein, a pioneer in the field of spatial inference from molecular networks, who presented to our community the exciting work he is doing in this field.
This autumn also marked the second iteration of the popular KTH master's/doctoral course Machine Learning for Biotechnology - co-taught with Anniina Vihervaara. Great to listen to the student presentations at the end!
Earlier this month, Ian traveled to Japan and gave a talk on spatial molecular networks at Kyoto University's Institute for Frontier Life and Medical Sciences (LiMe) hosted by Professor Hirofumi Shintaku.
And Ian is now on Bluesky - Follow and connect here!
Lastly, we recently admitted Krzysztof Mrosik to our ranks who will conduct his Master's thesis with us on the informatics and computational processing of information contained in DNA barcodes. Welcome Krzysztof!

Jul 3, 2024
Summer update - preprints, new member
This spring, we welcomed new member, experienced postdoctoral researcher Joel Spratt. Joel did his PhD in Oxford, and did a postdoc at Karolinska Institutet prior to joining us, where he will be working on developing new sequencing-based technologies. This spring we also published 2 preprints.
Check them out here:
Dahlberg SK, Bonet DF, Franzén L, Ståhl PL, Hoffecker IT. Hidden network preserved in Slide-tags data allows reference-free spatial reconstruction. bioRxiv. 2024:2024-06.
and here:
Bonet DF, Blumenthal JI, Lang S, Dahlberg SK, Hoffecker IT. Spatial coherence of DNA barcode networks. bioRxiv. 2024:2024-05.
Oct 10, 2023
Autumn update - new members, machine learning course,
🍂 Autumn greetings! 🍂 A warm welcome to two new lab members. Dr. Anastassia Runina is joining us as a postdoc. Anastassia, with background in real-time PCR, microarrays and next gen sequencing product development, and STI detection, will be developing novel biotechnologies related to next gen sequencing with us. Johanna Blumenthal, from the Molecular Techniques in the Life Sciences Scilifelab Master program, is joining us for her Project and Master's thesis. Welcome Anastassia and Johanna! In other news, Ian is in the final weeks of a new Master's course CB206V on Machine Learning in Biotechnology, co-organized with Gene Technology Department colleague Anniina Vihervaara. Check out the recorded lectures on Youtube.
Aug 8, 2023
End of summer news
Big news after an exciting spring and summer. We are honored to be one of only two groups in Sweden awarded an ERC Proof of Concept grant for our project MESH_CHIP, in June we filed our first patent, and this year Dr. Erik Benson, postdoc-extraordinaire, has accepted an offer to join Karolinska Institutet as an assistant professor next spring, right here in Scilifelab. Good times!!! 🔥🔥🔥
January 25, 2023
Happy new year
Happy new year and congratulations to Master's student Salomé Hahne who last week won a poster award in the KI Biomedicine program for her work with us on DNA diffusion analysis!🔥 And a belated congratulations to postdoc Erik Benson who was awarded the VR Starting Grant for his proposal on strong-binding DNA nanostructures! 💪 And finally a warm welcome to newcomer Hans Jiang, who is dual majoring in Biotechnology & Physics at KTH, and will be studying the thermodynamics and statistical mechanics of nucleic acid hybridization with us. Good start for the year!
November 16, 2022
New member & Ian teaching
Warm welcome to KI student Salomé Hahne! She has recently joined us as part of the Biomedicine Master's program's rotation. Ian taught a 2 day statistics and programming workshop for the Scilifelab Master's program this month. Video lectures can be found on youtube.
October 3rd, 2022
New DNA sequencing microscopy preprint
We're proud to announce our latest contribution to the emerging field of DNA sequencing-based microscopy. The work was led by newly enrolled doctoral student David Fernandez Bonet. Check out the full preprint.
August 1st, 2022 (retroactively posted)
Successful M.Sc. defenses
Congratulations to lab members Simon Kolmodin Dahlberg and David Fernandez Bonet on successful graduation and completion of their M.Sc. programs including successful defenses! We welcome them now as doctoral students. We wish you courage, strength, and good luck on your coming journey!
April 08, 2022
New molecular programming paper
New paper out on the stochastic modeling of antibody multivalent binding: nature.com/articles/s43588-022-00218-z
Check out our research briefing on the topic, and KTH news item for a reader-friendly explanation!
February 14, 2022
New members
We are welcoming three new members: Dr. Erik Benson has joined us for a postdoc after previously being at Oxford as an MSCA Fellow. Shuai Lang who was previously a Master's student at Karolinska Institutet will begin his doctoral studies with us. And KTH M.Sc. student Simon Kolmodin Dahlberg has joined us now for his thesis. Welcome!
November 17th, 2021
Welcoming new members
A warm welcome and thanks to our first generation of students. KTH students David Fernández Bonet and Simon Kolmodin Dahlberg will each do their Master's thesis with us. And a belated welcome to KI Master's student Li Ma who has worked over the summer and this autumn, establishing the first experiments and infrastructure of our new lab. Amazing job!
May 1st, 2021 (retroactively posted)
3, 2, 1...Launch!
The Hoffecker Lab has set up shop in the amazing Gene Technology department at KTH/SciLifeLab. It is a huge honor to be among this stellar group of internationally recognized researchers.
October 7th, 2020
Founding a new lab
The Hoffecker Lab will officially launch from May 1st 2021 as a new member in the Gene Technology Department, at KTH Royal Institute of Technology on the SciLifeLab campus in Stockholm. The funding making this possible comes from an ERC Starting grant , a starting grant from the Swedish Research Council (VR), co-financing from KTH's School of Engineering Sciences in Chemistry Biotechnology and Health (CBH), central funding from KTH, and a project grant from the Åke Wiberg Foundation.














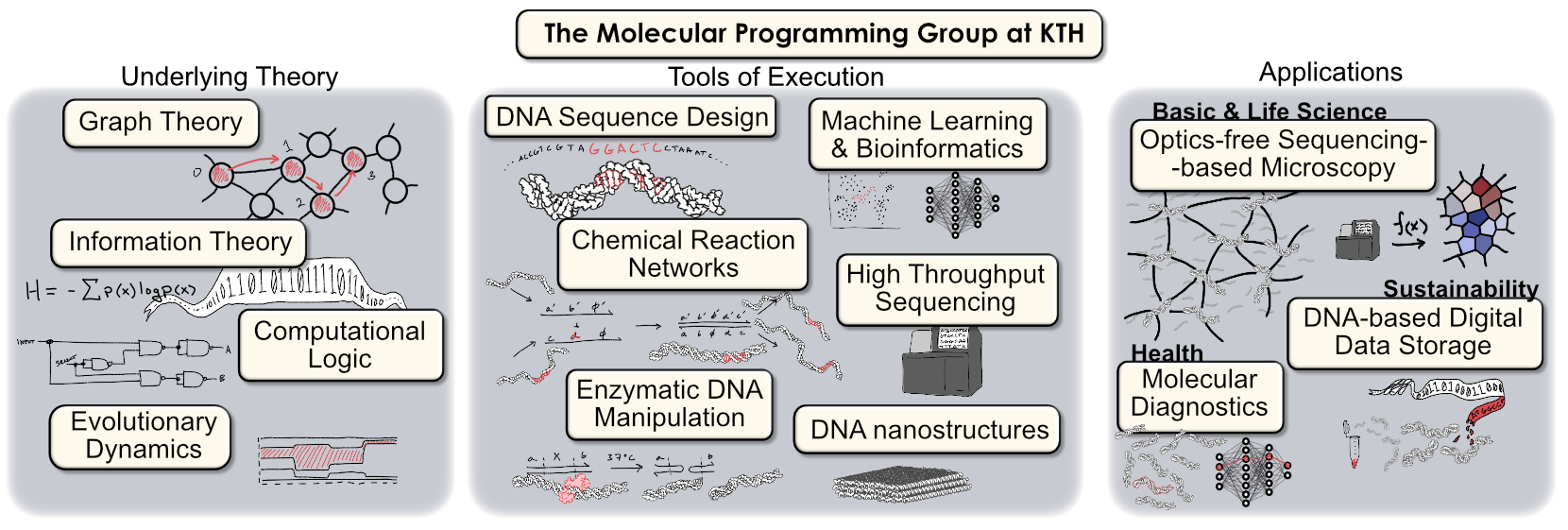
 The Molecular Programming Research Group is a team of scientist-engineers focused on the development of new biotechnologies using DNA as an engineering material. We are part of the Science for Life Laboratory (SciLifeLab) and KTH Royal Institute of Technology in Stockholm, Sweden. Our group is led by Dr. Ian Hoffecker, a specialist in polymer engineering, DNA nanotechnology, and the physics of computation.
DNA is not only the carrier of genetic information in living organisms, but also a programmable polymer that can be artificially synthesized with unique properties that make it an ideal material for engineering at the nanoscale. By exploiting the predictable base-pairing interactions of DNA, we can design and construct complex molecular systems that perform specific functions, opening up new possibilities in biotechnology, medicine, and materials science.
These days, sequencing DNA is faster and cheaper than ever before, enabling new ways to read out information encoded in DNA molecules. Our thesis is simple: so long as humans inhabit biological bodies, DNA will remain a relevant molecular information processing vehicle, and the ecoysystem of tools for analyzing and manipulating it will continue to grow in power and accessibility.
By treating DNA as a programmable material, we can create new capabilities for sensing, analyzing, and manipulating nanoscale, biological, and even artificial systems.
The Molecular Programming Research Group is a team of scientist-engineers focused on the development of new biotechnologies using DNA as an engineering material. We are part of the Science for Life Laboratory (SciLifeLab) and KTH Royal Institute of Technology in Stockholm, Sweden. Our group is led by Dr. Ian Hoffecker, a specialist in polymer engineering, DNA nanotechnology, and the physics of computation.
DNA is not only the carrier of genetic information in living organisms, but also a programmable polymer that can be artificially synthesized with unique properties that make it an ideal material for engineering at the nanoscale. By exploiting the predictable base-pairing interactions of DNA, we can design and construct complex molecular systems that perform specific functions, opening up new possibilities in biotechnology, medicine, and materials science.
These days, sequencing DNA is faster and cheaper than ever before, enabling new ways to read out information encoded in DNA molecules. Our thesis is simple: so long as humans inhabit biological bodies, DNA will remain a relevant molecular information processing vehicle, and the ecoysystem of tools for analyzing and manipulating it will continue to grow in power and accessibility.
By treating DNA as a programmable material, we can create new capabilities for sensing, analyzing, and manipulating nanoscale, biological, and even artificial systems.